Novel Species Research
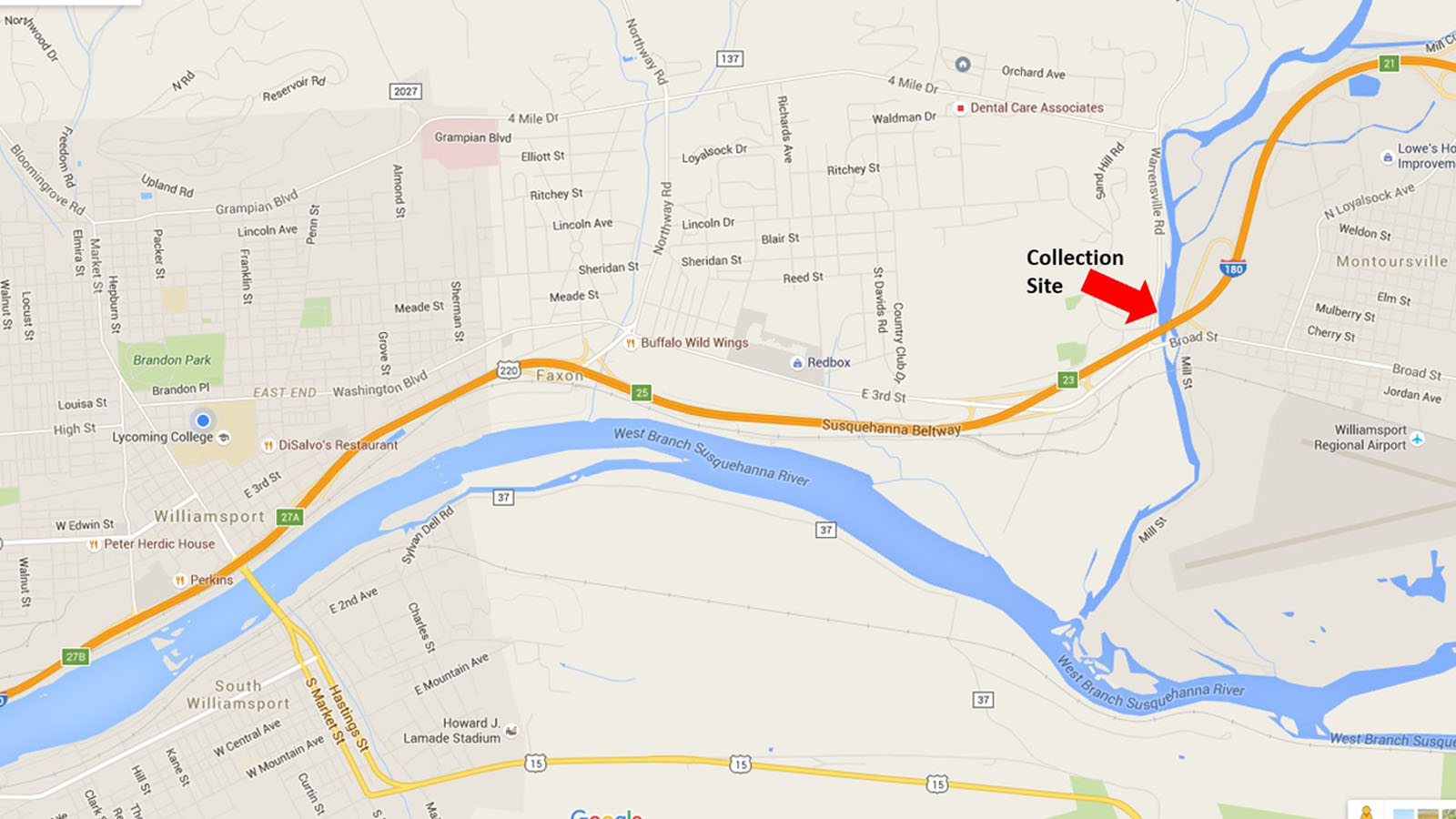
Research in the Newman Lab begins in the Bio321 General Microbiology course. The description below corresponds to our current pipeline, which has evolved over the past 10 years or so. In the LycoMicro course, we identify unknown organisms isolated from the Loyalsock Creek, near the Broad Street and Route 220/I-180 bridges into Montoursville.
 |
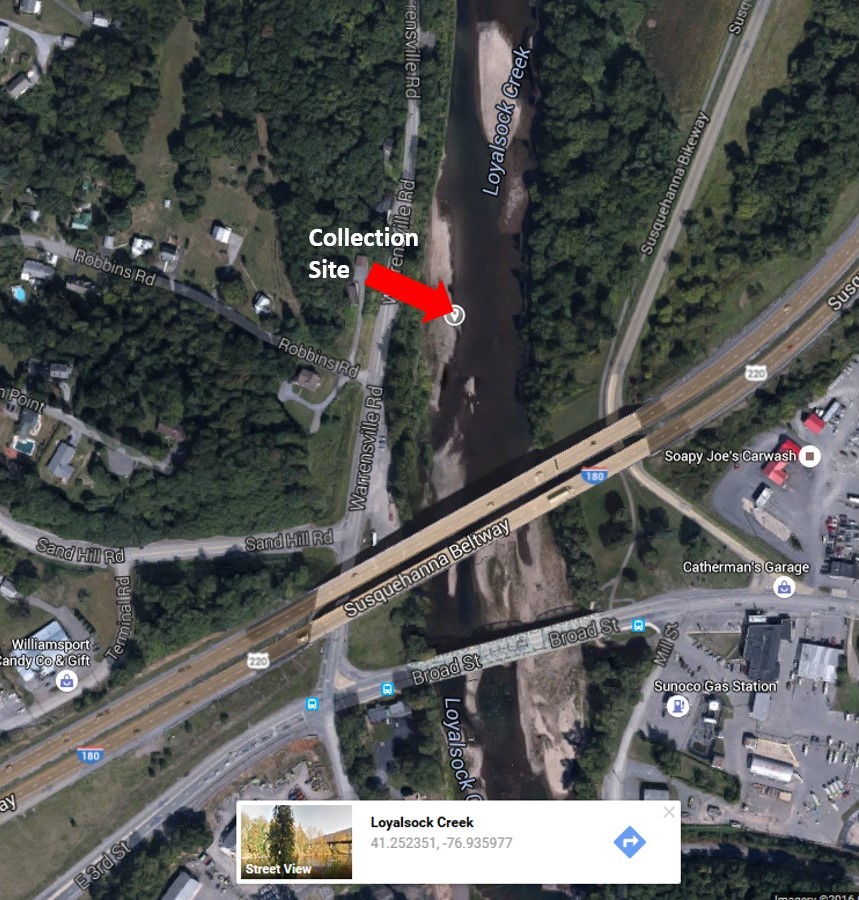 |
Because the course is taught every Spring semester, samples are usually collected between Christmas & New Year’s Day and spread onto TSA and R2A plates. After 2 days growth at room temperature (22oC), well isolated colonies are picked and patched onto 2 replica plates, one incubated at 37oC, the other at 22oC. Patches that grow at 22oC, but NOT at 37oC, are then restreaked onto R2A and provided to the students as unknown organisms. Pre-screening to remove organisms that grow at 37oC minimizes the possibility of obtaining human pathogens.
Please visit the Bio321 course website for detailed information on the characterization done in the course. We frequently obtain organisms whose 16S rRNA sequence is less than 98.5% identical to any validly published & named species. We consider these strains to be novel species candidates, and students often will follow-up on their organism (or one of the many stored in the -80oC freezer) as part of the Bio447 Research Methods course or as an Independent Study project.
Our goal is to characterize, publish and name these novel species. In the process, we have recognized what a mess microbial taxonomy is, and hope to improve the system by providing procedures on what we consider to be best practices. We’d also like to use the expertise we have developed to help others characterize their novel organisms. These are really fantastic projects for undergraduates! Feel free to get in touch if we can be of assistance. Please be patient as I (Jeff Newman) have a very heavy teaching and service load.
We are grateful to the NSF for a Major Research Instrumentation grant (MRI) obtained in 2009 to purchase:
We have also been involved with the Genome Consortium for Active Teaching – NextGen Sequencing Group (GCAT-SEEK) which has been funded by the NSF and HHMI to provide access to NextGen sequencing for undergraduate education.
NewmanLab Novel Microbe Publication Pipeline
1. 16S Sequence Deposit
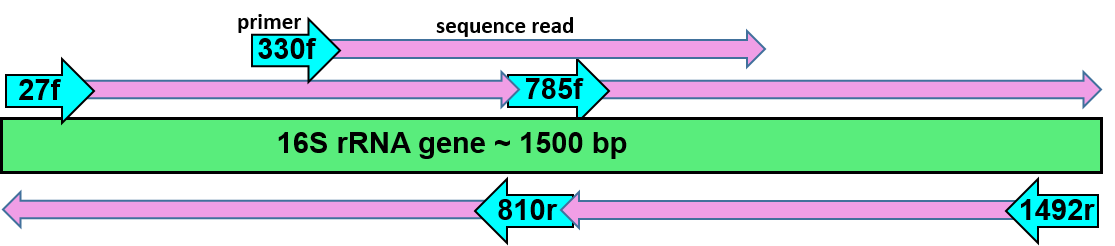
If 16S sequence with primer 27f from Microbiology course is less than 98.5% identical to other validly published species using EzTaxon, obtain nearly complete 16S sequence using primers 330f, 785f, 1492r, 810r. Assemble with CAP3, deposit in NCBI, obtain accession.
2. 16S Tree & Matrix
Use EzTaxon to identify, perform Comp Gen analysis with closest Genus, download sequences (Fasta zZ), replace accessions, add outgroup (type species from next closest genus), align with MEGA, Create NJ, MP, ML trees (save .mts and caption). Create publication quality NJ tree, determine closest relatives for comparative analysis.
3. Culture Collections
Obtain closest relatives from culture collection, deposit organism into at least two culture collections. Obtain Deposit Certificates.
| Abbreviation | Name | Notes |
| DSMZ | German Collection of Microorganisms and Cell Cultures | Catalogue of microorganisms |
| BCCM | Belgian Co-ordinated Collections of Micro-organisms | On-Line Catalogues BCCM/LMG bacteria catalogue |
| LMG | Laboratorium voor Microbiologie, Universiteit Gent, Belgium | Bacteria Catalogue |
| KACC | Korean Agricultural Culture Collection | |
| KCTC | Korean Collection for Type Cultures | Online Catalogue |
| CCUG | Culture Collection, University of Göteborg, Sweden | Search CCUG databases Type strains |
| JCM | Japan Collection of Microorganisms | JCM On-line Catalogue |
| ACM | Australian Collection of Microorganisms | Search ACM |
| ARS | Agricultural Research Service Culture Collection, USA | Search the Collection |
| ATCC | American Type Culture Collection, USA | Search facilities available |
| CECT | Colección Española de Cultivos Tipo (Spanish Type Culture Collection) | Search bacteria |
| CIP | Centre de Ressources Biologiques de l'Institut Pasteur (English Version) | Online catalogue |
| MTCC | Microbial Type Culture Collection & Gene Bank (MTCC), India | Search facilities available |
| NBRC | NITE (National Institute of Technology and Evaluation) Biological Resource Center, Japan | NBRC Culture Catalogue Search |
| NCTC | National Collection of Type Cultures, UK | Bacteria, Plasmids, Mycoplasmas, Plasmids & Transposons Search |
| VKM | All-Russian Collection of Microorganisms | VKM Catalogue |
| CGMCC | China General Microbiological Culture Collection Center |
4. Sequence Genome(s)
Isolate genomic DNA from novel and reference strains, PCR amplify 16S rRNA, confirm correct DNA by Sanger sequencing, sequence genome(s) with MiSeq (2 x 300). Annotate with RAST, obtain GC%.
5. Identify Closest Genomes
Determine estimated DNA-DNA hybridization (GGDC eDDH), Average Amino Acid Identity (AAI), values for novel vs all genomes in the genus and create a table. Some genomes sequenced by JGI-IMG may not have been deposited in GenBank yet, therefore It may be necessary to download genomes (*.gbk preferably, or *.fasta files) from JGI-IMG, upload them to RAST for annotation, then, when annotation is complete, download the contig fasta file from RAST to use for GGDC calculation.
6. Phylogenomic Matrices
Create matrix for eDDH & AAI with novel, top 8 most similar type strains in genus, plus type species of genus (see List of Prokaryotic Names with Standing in the Nomenclature - LPSN), and type species of next closest genus (see Living Tree Project – LTP). Create matrix with Reciprocal Orthology Score Average (ROSA) and 16S seq for same organisms, Average Nucleotide Identity (ANI) list for novel vs same organisms. Select and obtain additional organisms for phenotypic comparison if needed.
If the most similar genome has an AAI below 70, it is likely that they are separate genera, and you may have a new genus! There currently is no firm threshold for what constitutes a new genus, therefore, it is helpful to demonstrate the range of AAI values between different genera in the family (-aceae) of your organism.
If you think your organism may represent a new genus, visit the LPSN, search for the family name and click on the link for a list of all genera in the family. For each genus, download a genome from NCBI or IMG as above, and upload to RAST. Ideally, you should use the type strain of the type species (specified in LPSN). For each genus, use EzTaxon to compare the type strain of the type species to the database to see which other genera are most closely related so that they can be grouped for creation of AAI matrices as described above.
7. Venn Diagram
Use RAST Sequence-based Comparison Tool to compare novel, top 3 most similar type strains in genus, plus type species of genus. Create Venn Diagram.
Venn Diagram Calculator Files developed by Tom Sontag'13 & Andrew Gale'15. Please contact Jeff Newman to request use of calculator files because we have not yet published this work.
8. Unique, Shared Genes
Create list of unique genes in novel organism, or interesting sets of shared genes.
9. Obtain & Read Primary and Secondary Literature
Papers about the Phylum, Family, Genus, species to be compared, Minimal standards, Bergey’s Manuals of Determinative and Systematic Bacteriology.
10. Phenotypic Comparison of Novel with Type Strains of Closest Relatives
a. Growth at different temperatures, pH, [NaCl], oxygen
b. Gram Stain, Wet Mount, Endospore Stain, Acid Fast Stain
c. Kirby-Bauer for antibiotic sensitivity
d. Traditional Carbon, Nitrogen metabolism tests
e. Exoenzymes, Differential and selective medium
f. Biolog GenIII Plates
g. API Test Strips
h. Fatty Acid Methyl Ester (FAME) analysis by Gas Chromatography (GC)
i. Polar Lipid Analysis by Thin Layer Chromatography (TLC)
j. Respiratory Quinones by TLC & HPLC
k. Other Family or Genus-specific tests
11. Compare results obtained with closest relatives to published results, genome annotation.
12. Prepare table of differentiating characteristics for publication.


